Bihar Board Class 12 Biology Solutions Chapter 6 Molecular Basis of Inheritance Textbook Questions and Answers, Additional Important Questions, Notes.
BSEB Bihar Board Class 12 Biology Solutions Chapter 6 Molecular Basis of Inheritance
Bihar Board Class 12 Biology Molecular Basis of Inheritance Text Book Questions and Answers
Question 1.
Group the following as nitrogenous bases and nucleosides :
Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine.
Answer:
Among the above Adenine, Thymine, Uracil and Cytosine are nitrogenous bases whereas Cytidine and Guanosine are nucleosides. Nucleosides are formed by a purine or pyrimidine nitrogenous base and pentose sugar linked together through a N-glycosidic linkage.
Question 2.
If a double-stranded DNA has 20 per cent of cytosine, calculate the per cent of adenine in the DNA.
Answer:
Assuming that the total number of nitrogenous bases is 100 in the DNA sequence. Out of this Cytosine is 20 in number. According to the base-pairing rule, we know that in a DNA segment a cytosine always pairs up with a guanine by three hydrogen bonds. Thus the number of guanine is also 20. Now, we are left with 60 base pairs. Adenine always pairs up with Thymine by two hydrogen bonds. So equal number of Adenine and Thymine must be present in the sequence, thus the amount of Adenine would be (60+2=30) 30%.
Cytosine is 20%
Cytosine = Guanine So, 20% = 20%
Total 40%
number of Adenine and Guanine is 100 – 40 = 60%
Adenine = Guanine
So, 60% divided in equal amounts 30% + 30%
So, Adenine amount would be 30%.
Question 3.
If the sequence of one strand of DNA is written as follows :
5-ATGCATGCATGCATGCATGCATGCATGC-3′
Write down the sequence of complementary strand in 5’→3′ direction.
Answer:
In DNA, the two strands are complementary to each other i.e. the other strand will pair up according to the sequence in the first strand. Every time Adenine pairs with Thymine by two hydrogen bonds and Cytosine pairs with Guanine by forming three hydrogen bonds. So, it’s easy to predict the sequence of other strand if the base pair sequence of one strand is known. Here, the sequence of the complementary strand would be :
5′-ATGCATGCATGCATGCATGCATGCATGC-3′
3’-TACGTACGTACGTACGTACGTACGTACG-5′ (Complementary strand)
Question 4.
If the sequence of Coding strand in a transcription unit is written as follows :
5’-ATGCATGCATGCATGCATGCATGCATGC-3′
Write down the sequence of mRNA.
Answer:
Coding strand has a polarity 5’ →3′ direction and it does not code for RNA because the enzyme RNA polymerase (RNA polymerisation enzyme) can catalyze RNA polymerization only in one direction i.e. 5’→3′. Thus the 3′ polarity stand can only act as the template for transcription. That is called as template strand.
We can also conclude the sequence of mRNA from the coding strand. It would be same as the coding strand except, there would be Uracil opposite to the Adenine in place of Thymine. In case of template strand the mRNA sequence would be complementary to the template strand just replacing Thymine by Uracil.
Here, the mRNA sequence would be :
5VATGCATGCATGCATGCATGCATGCATGC-3′ Coding strand
5-AUGCAUGCAUGCAUGCAUGCAUGCAUGC-3′ (mRNA sequence).
![]()
Question 5.
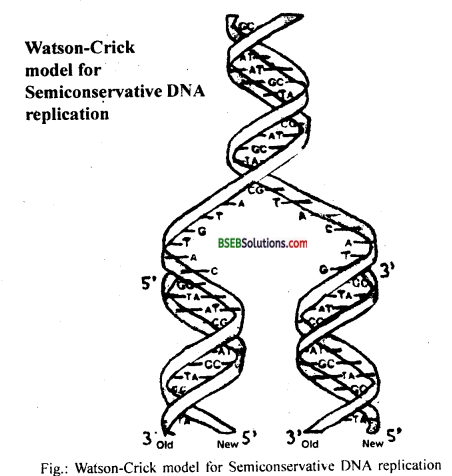
Which property of DNA double helix led Watson and Crick to hypothesise semiconservative mode of DNA replication? Explain.
Answer:
DNA is the predominant genetic material. It has got several properties to suggest its role as genetic material. It can replicate; it is chemically and structurally stable and there is scope for mutations and thus for evolution. Watson and Crick purposed the double-helical structure of DNA, they .have also suggested the semiconservative model for DNA replication. During replication, DNA strands uncoil and both the strands act as a template for the synthesis of new complementary strand.
Because of the rule of base pairing and complementarity, DNA has the ability to direct its duplication. Thus at the end of replication, each daughter DNA molecule would have one parental strand and one newly synthesized strand. This mode of replication is termed as semiconservative replication.
Question 6.
Depending upon the chemical nature of Template (DNA or RNA) and the nature of nucleic acids synthesised from it (DNA or RNA), list the types of nucleic acid polymerases.
Answer:
There are two types of nucleic acids. Deoxyribonucleic acid and ribonucleic acid. These are synthesized within the cellular
machinery by specific enzymes known as polymerases. Polymerases are of two types depending upon the kind of nucleic acid.
(i) DNA polymerase: It is DNA dependent enzyme, along with many additional enzymes, it uses a DNA template to catalyze the polymerization of deoxynucleotides. DNA polymerase can catalyze polymerization only in one direction i.e. 5′-3′ direction. Due to this, replication on strand (3′-5′ direction) is continuous but on the other (5’→3′ direction) it is discontinuous or takes place in fragments.
(ii) RNA polymerase: These are the enzymes which catalyse the polymerization of RNA. These may be different types for e.g. in bacteria there are three types of RNAs : messenger RNA (mRNA), transfer RNA (tRNA) and ribosomal RNA (rRNA). All three types of RNAs are synthesized by single DNA-dependent RNA polymerase. It catalyses transcription of all three types of RNA in bacteria, which are needed in protein synthesis.
In eukaryotes, there are three RNA polymerases present in the nucleus and additional RNA polymerases are present in organelles. RNA polymerase I does transcription of rRNAs (28s, 18s, 5.8s); RNA polymerase II transcribes precursor of mRNA, hnRNA (heterogeneous nuclear RNA); RNA polymerase III does transcription of tRNA, 5sRNA and snRNAs (small nuclear RNAs).
Question 7.
How did Hershey and Chase differentiate between DNA and protein in their experiment while proving that DNA is the genetic material?
Answer:
In 1952, Alfred Hershey and Martha Chase worked to discover and prove that DNA is the genetic material and not the proteins. They worked with bacteriophage T,, which attacks bacterium E.Coli.
Hershey and Chase (1952) conducted their experiment on T, phage which attacks the bacterium Escherichia coli. The phage particles were prepared by using radioisotopes of 35S and ’:P in the following steps :
(i) Few bacteriophages were grown in bacteria containing ’5S. This radioactive 35S was incorporated into the cysteine and methionine amino acids of proteins and thus these amino acids with 35S formed the proteins of phage.
(ii) Some other bacteriophages were grown in bacteria having,2P. This radioactive -2P was restricted to DNA of phage particles.
These two radioactive phage preparations (one with radioactive proteins and another with radioactive DNA) were allowed to infect the culture of E.coli. The protein coats were separated from the bacterial cell walls by shaking and centrifugation.
The heavier infected bacterial cells during centrifugation pelleted to bottom (fig.). The supernatant had the lighter phage particles and other components that failed to infect bacteria. It was observed that bacteriophages with radioactive DNA gave rise to radioactive pellets with 32P in DNA.
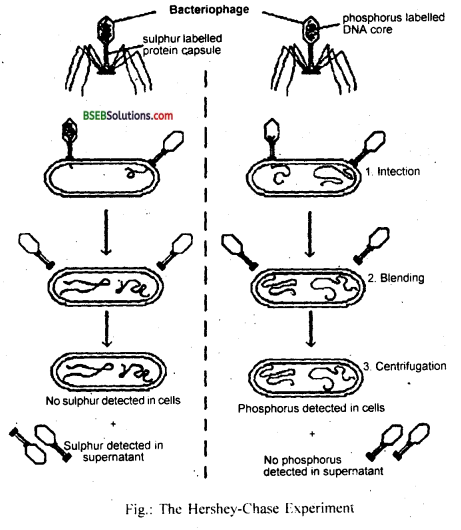
In the phage particles with radioactive protein, the bacterial pallets have almost nil radioactivity indicating that proteins have failed to migrate into bacterial cell. So it was concluded that DNA must have entered the bacteria.
Question 8.
Differentiate between the followings :
(i) Repetitive DNA and Satellite DNA
(ii) mRNA and small tRNA
(iii) Template stand and coding stand
Answer:
Difference between Repetitive DNA and Satellite DNA :
(i) Repetitive DNA is DNA fingerprinting which involves identifying difference in some specific regions in DNA sequences. Satellite DNA is the bulk DNA which forms a major peak and the other small peaks.
(ii) Difference between mRNA and small tRNA: mRNA sequence decides the sequence of amino acids tRNA acts as an adopter molecule.
(iii) Difference between Template Strand and Coding Strand: Template Strand – The two strands that have opposite polarity and the DNA-dependent RNA polymerase also catalyse the polymerization in only one direction, i.e:, 5-3′ the strand that has the polarity 3′-5′ acts as a template and is referred as template strand. Coding strand – The strand which has the polarity 5′ – 3′ and the sequence same as RNA (except thymine at the place of uracil) is displaced during transcription. This strand is referred to as Coding strand.
![]()
Question 9.
List two essential roles for ribosome during translation.
Answer:
Translation is the process of polymerization of amino acids to form polypeptides. Ribosome and ribosomal RNA play important role during translation. We can say ribosome is the cellular factory responsible for synthesizing proteins. Ribosome consists of structural RNAs and arround 80 different proteins. It consists of two subunits: a large subunit and a small subunit. For initiation of translation the ribosome, binds to the mRNA at the start codon.
Then the ribosome proceeds to the elongation phase by adding amino acids to the polypeptide chain. During this phase the ribosome moves from codon to codon along the mRNA. Amino acids are added one by one. They are translated into polypeptide sequences dictated by DNA and represented by mRNA. At the end, on encounter of stop condon the process of translation terminates and both the subunits of ribosome detach. Thus the ribosome serve the essential functions in the process of translation and it also acts as a catalyst for the formation of peptide bond.
Question 10.
In the medium where E.coli was growing, lactose was added, which induced the lac-operon. Then, why does lac-operon shut down after some time after addition of lactose in the medium?
Answer:
When a polycistronic structural gene is regulated by a common promoter and regulatory genes, the whole unit is called an operon, e.g. lac operon. It was explained by Francis Jacon and Jacque Monod (1961).
It was found that when sugar lactose is added to the E.coli, cultures it induces synthesis of three enzymes required for degradation of lactose into glucose and galactose. These are P-galactosidase, permease and transacetylase. The process is called as induction and the enzymes as indupible enzymes.
In lac operon, when lactose is added, it enters the cell with the help of permease, a small amount of which is already present in cell. Lactose binds itself to active repressor and changes its structure. The repressor now fails to bind to the operator. Then RNA polymerase starts transcription of operon by binding to promoter site P. All the three enzymes for lactose metabolism are synthesized. Finally all the lactose molecules are used up. The whole process of induction can be understood by following figure.
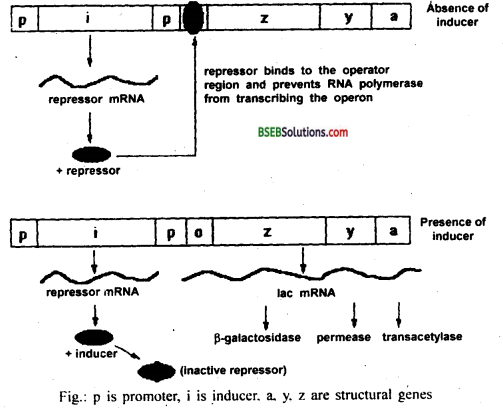
After some time when whole of lactose is consumed, there is no inducer (lactose) present to bind to the repressor. Then the repressor becomes active again, attaches itself to the operator and finally switches off the operon.
Question 11.
Explain (in one or two lines) the function of following :
(a) Promoter,
(b) tRNA,
(c) Exons.
Answer:
(a) Promoter: Promoter gene marks the site at which transcription of mRNA starts and where RNA polymerase enzyme binds. The promoter is located towards 5′-end (upstream) of the structural gene. The presence of promoter sequence in a transcription unit defines the template and coding strands.
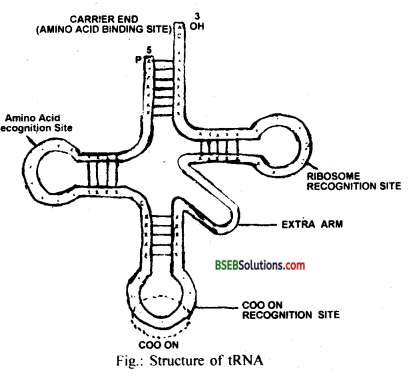
(b) tRNA: It is transfer RNA, helps in the process of protein synthesis. tRNA brings the aminoacids and reads the genetic code. It contains ribosome recognition site, anticodon site, amino acid activating synthetase enzyme recognition site (DHU arm) and carrier end which acts as amino acid binding site. tRNa is single-stranded polymer of less than one hundred subunits. It is present in the cytoplasm.
(c) Exons: Exons are those sequences which appear in mature or processed RNA. In eukaryotes, the monocistronic structural genes have interrupted coding sequences or the eukaryotic genes are split genes. The coding sequences or expressed sequences are called as exons. These exons are interrupted by introns or intervening sequences. Introns are those sequences which do not code for mRNA. These sequences do not appear in mature or processed
RNA.
Question 12.
Why is the human genome project called a megha project?
Answer:
Human Genome Project (HGP) is called a megha project became of the following reasons :
Human genome is said to have approximately 3×109bp, and if the cost of sequencing required is US $3 per bp, the total estimated cost of the project would be approximately 9 billion US dollars.
If the obtained sequences were to be stored in typed form in books, and if each page of the book contained 1000 letters and each book contained 1000 pages then 3300 such books would be required to store the information of DNA sequence from a single human cell. The enormous amount of data expected to be generated also required the use of high-speed computational devices for data storage and analysis.
HGP is also closely associated with the rapid development of a new area in biology called as Bioinformatics The project was completed in 2003. Knowledge about of the effects of DNA variations among individuals can lead to revolutionary new ways to diagnose, treat and present* thousands of disorders that affect human beings.
Learning about non-human organisms, DNA sequences can lead to an understanding of their natural capabilities that can be applied toward solving challenges in health care, agriculture, energy production, environmental remediation, etc.
Some Important goals of HGP :
- Identify all the 20,000 – 25,000 genes in human DNA.
- To determine the sequences of the 3 billion chemical base pairs that make human DNA.
- Store the information in databases.
- Improve tools for data analysis.
- Transfer related technologies to other sectors, such as industries, etc.
- Address the ethical, legal and social issues (ELS1) that may arise from the project.
Question 13.
What is DNA fingerprinting? Mention its application.
Answer:
if one aims to final out genetic differences between two individuals or among individuals of a population, sequencing the DNA every time would be a daunting and expensive. DNA fingerprinting is a very quick way to compare the DNA sequences of any two individuals.
Application of DNA fingerprinting: In addition to application in forensic science, it has much wider application, such as in determining population and genetic diversities.
![]()
Question 14.
Describe briefly the followings :
(a) Transcriptions
(b) Polymorphism
(c) Translation
(d) Bioinformatics
Answer:
(a) Transcription – ft is a process of copying genetic information from one strand of the DNA into RNA is termed as transcription.
(b) Polymorphism – It refers to variation of genetic level. This information revolutionise the processes of finding chromosomal locations for disease-associated sequences and tracing human history.
(c) Translation – It is a process that requires transfer of genetic information from a polymer of neuleolides to a polymer of amino acids to form a polypeptide.
(d) Bioinformatics – A new area in biology is reformed to as bioinformatics. Knowledge from the DMA sequence, will help research through the coming decides leading to the understanding of biological system.
Bihar Board Class 12 Biology Molecular Basis of Inheritance Additional Important Questions and Answers
Very Short Answer Type Questions
Question 1.
Who coined the word gene?
Answer:
Johannsen coined the word gene.
Question 2.
What is genetic material?
Answer:
DNA and RNA in some bacteria.
Question 3.
What does DNA and RNA stands for?
Answer:
DNA: deoxyribonucleic acid; RNA: ribonucleic acid.
Question 4.
What are purines and pyrimidines?
Answer:
These are the nitrogenous bases present in the nucleic acids. There are two purines viz. Adenine and Guanine and three pyrimidines viz. Thymine, Cytosine and Uracil.
![]()
Question 5.
What is the haploid content of human DNA?
Answer:
3.3 × 109 bp.
Question 6.
What does DNA’s backbone made of?
Answer:
DNA’s backbone is made up of repetitive sugar and phosphate moieties.
Question 7.
Name the scientist who identified DNA.
Answer:
Friedrich Meischer (1869) first identified DNA and named it nuclein.
Question 8.
What is the polarity of DNA strands?
Answer:
Two strands of DNA have antiparallel polarity i.e. if one strand has 5’→3′ polarity, the other will have 3’→5′ polarity.
Question 9.
What is the rule of base pairing?
Answer:
It states that each time Adenine will form two hydrogen bonds with Thymine (Uracil in case of RNA A=U) and Guanine will bind to Cytosine by three hydrogen bonds.
Question 10.
Write down central Dogma.
Answer:
![]()
Question 11.
What are histones?
Answer:
Histones are positively charged basic proteins. They assist in wrapping arround the negatively charge DNA.
Question 12.
What are bacteriophages?
Answer:
Bacteriophages are viruses that infect bacteria.
![]()
Question 13.
Which was the first genetic material?
Answer:
RNA was the first genetic material.
Question 14.
What do you understand by discontinuous replication?
Answer:
The replication of DNA stand which have a polarity of 5’→3′ direction is discontinuous. It takes place in small fragment by DNA polymerase.
Question 15.
Define transcription.
Answer:
Transcription is the process of copying genetic information from one DNA strand to RNA.
Question 16.
What are the components of transcription unit?
Answer:
A promoter; the structural gene and a terminator.
Question 17.
Define a gene.
Answer:
A gene is a functional unit of inheritance. It may also be said as a DNA sequence coding for tRNA or rRNA molecule.
Question 18.
What do you mean by degeneracy of genetic code?
Answer:
The genetic code is degenerate, it means that one amino acid can be coded by more than one codon.
Question 19.
What is translation?
Answer:
Translation is the process of polymerization of amino acids to form a polypeptide, whose sequence is defined by mRNA.
![]()
Question 20.
What is lac-operon?
Answer:
It is the operon present in bacteria which codes for genes responsible for metabolism of lactose.
Short Answer Type Questions
Question 1.
Why is the DNA molecule compared to a spiralling staircase? Which three components make up the nucleotide?
Answer:
DNA is made up of two polynucleotide chains, which are helically coiled so that the nitrogenous bases of two strands are linked together by hydrogen bonds. The two right-handed helices are coiled in an interlocked form about the same axis. The DNA strands are made up of repeating nucleotide units. A nucleotide is made up of purine or pyrimidine nitrogenous bases which are joined by deoxyribose sugar, which is linked with phosphate group.
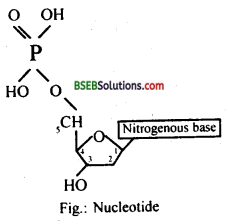
Question 2.
What are three types of RNA molecules? How is each related to the concept of information flow?
Answer:
There are three major types of RNAs: messenger RNA (mRNA), tRNA (transfer RNA) and ribosomal RNA (rRNA). These three are required for the process of protein synthesis (translation). The mRNA provides the template for protein synthesis, tRNA brings amino acids and reads the genetic code and rRNA plays structural and catalytic role during translation.
Question 3.
What is a peptide bond? How is it formed?
Answer:
In a polypeptide chain, the adjust amino acids are linked to each other by a peptide bond.
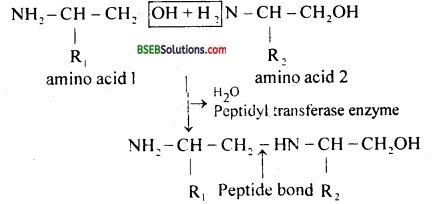
Question 4.
What is the role of nonsense codon in protein synthesis?
Answer:
Nonsense codon as the name indicates, code for nothing in a growing polypeptide chain thus the addition of new amino acid stops and the process of translation stops. Thus nonsense codons act as stop signals and terminate the translation process. There are three nonsense condons viz. UAG, LAA and UGA.
![]()
Question 5.
What are three essential requirements of the genetic material?
Answer:
Genetic material must fulfil the following criteria :
- It should be able to replicate.
- It should be chemically and structurally stable.
- It should provide the scope for mutation and evolution.
Question 6.
Differentiate between DNA and RNA.
Answer:
Differences between DNA and RNA :
| DNA | RNA |
| (1) It usually occurs inside nucleus and some cell organelles. | (1) Very little RNA occurs inside nucleus. Most of it is found in the cytoplasm. |
| (2) DNA is the genetic material. | (2) RNA is not the genetic material except in certain viruses, e.g. Reovirus. |
| (3) It is double-stranded with the exception of some | (3) RNA is single-stranded with the exception of viruses (e.g. Φ × 174). |
| (4) DNA contains over a million nucleotides. | (4) Depending upon the type, RNA contains 70-12000 nucleotides. |
| (5) Molecular weight ranges from 3-4 million in Escherichia coli to 263 million in chromosome l of human beings. | (5) Molecular weight ranges from 25000-2,000,000. |
| (6) It is Fuelgen positive. | (6) RNA is Fuelgen negative |
| (7) DNA is of only two types, intra-nuclear and extra-nuclear. | (7) There are at least three types of RNAs – rRNA rRNA and tRNA. |
| (8) It contains deoxyribose sugar. | (8) It contains ribose sugar. |
| (9) Nitrogen base thymine occurs in DNA along with three others – adenine, cytosine and guanine. | (9)Thymine is replaced by uracil in RNA. The other three are similar adenine cytosine and guanine. |
| (10) Unusual bases are very few or absent. | (10) Many unusual or modified bases are often present. |
| (11) Renaturation after melting is slow. | (11) It is quite fast. |
| (12) Hydrogen bonds are formed between complementary nitrogen bases of the opposite strands of DNA (A – T, C – G). | (12) Base pairing through hydrogen bonds occurs only in the coiled parts. |
| (13) DNA is spirally twisted to produce a regular helix. | (13) The strand may get folded at places to produce a secondary helix or pseudo helix. |
| (14) It replicates to form new DNA molecules. | (14) It cannot normally replicate itself. |
| (15) DNA replication requires a primer some viruses (e.g., double-stranded in Reovirus). | (15) No primer is needed during the formation of RNA. |
| (16) DNA transcribes genetic, information to RNA. | (16) RNA translates the transcribed message for forming polypeptides. |
| (17) Its quantity is fixed for cells. | (17) The quantity of RNA of a cell is variable. |
| (18) DNA controls metabolism and genetics including variations. | (18) ftoniy controls metabolism under instructions from DNA. |
| (19) Purine and pyrimidine base are in equal number. | (19) There is no proportionality between number of purines and pyrmidine bases. |
| (20) It occurs in the form of prochromosome, chromatin or chromosomes. | (20) It occurs in ribosomes or forms association with ribosomes. |
| (21) 3H precursor is 3H-thymidine. | (21) H precuror is 3H-uridine. |
| (22) It is long-lived. | (22) Some RNAs are very short-lived while others have somewhat longer life. |
| (23) It can be hydrolysed by DNA-ase. | (23) It is hydrolysed by RNA-ase. |
Question 7.
Enlist different types of DNA.
Answer:
(i) A-DNA: It has i l base pairs, which are tilted from the axis of the helix (γ = 20.2”). It is found at 75% relatine humidity in the presence of Na+, K+ or Cs+ ions.
(ii) B-DNA: It has 10 base pairs which are tilted from the helix axis (γ = 6.3°). It is found at 92% relative humidity and low ionic strength.
(iii) C-DNA: It has 9.33 base pairs which is negatively tilted from helix axis (γ = 7.8°). It is found at 66% relative humidity in the presence of lithium.
(iv) D-DNA: It has 8 base pairs, negatively tilted from the helix axis (γ = 16.7°).
(v) Z-DNA : It is left-handed double helix DNA with a zig-zag sugar-phosphate backbone in antiparallel manner. It has 12 base pairs, which are tilted from helix axis (γ = 7°).
![]()
Question 8.
What are the characteristics of genetic code?
Answer:
- The codons are tripled.
- One codon codes for only one amino acid. So genetic code is specific and unambiguous (clear).
- Genetic code is degenerate i.e. some amino acids are coded by more than one codon.
- The codon is read in a contiguous manner, without any punctuations.
- The codon is universal.
Question 9.
What is the purpose of proofreading in DNA synthesis?
Answer:
The process of replication is efficient and accurate. Specificity of base-pairing ensures exact replication. Sometimes if any wrong base is present (l in 10,000), it can be removed by the proof-reading activity. It is done by the enzyme DNA-polymerase.
Question 10.
What are important features of DNA structure?
Answer:
- DNA, double helix is made up of two polynucleotide chains. The backbone is made up of sugar-phosphate and the bases project inside.
- The two strands are antiparallel i.e. one with 5′ →3′ polarity and the other with 3’→5′ polarity;
- The bases pair with each other via hydrogen bonds. Adenine forms two hydrogen bonds with thymine and guanine pairs with cytosine by three hydrogen bonds.
- The two strands are coiled in right-handed fashion. One turn contains 10 base pairs.
Question 11.
Which molecule bears codons and which molecule anticodons?
Answer:
mRNA bears codons, it acts as the reading frame. mRNA codes information in the form of codons for the synthesis of proteins. One codon consists of three nucleotides, it codes for a specific amino acid on a growing polypeptide chain. tRNA bears anticodons on its anticodon loop which is having complementary bases to the codon present on mRNA.
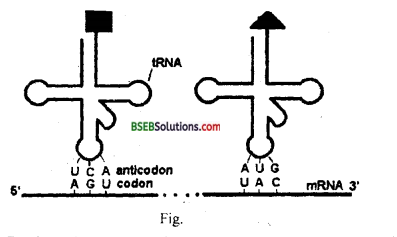
Question 12.
Briefly describe the process of replication.
Answer:
The process of replication starts from unwinding of two strands, which will act as a template for the synthesis of new complementary strands. DNA dependent DNA polymerase enzyme catalyzes the polymerization of deoxynucleotides. Replication occurs at small opening of the DNA helix called as replication fork. DNA polymerase can catalyze polymerization only in one direction i.e. 5’→3′ which is called continuous replication. The other strand replicated by discontinuous mechanism. So the continuous strand is with polarity (3′ →5′) and discontinuous strand is with poliarity (5’→3′). The discontinuous fragments are joined by DNA ligase enzyme.
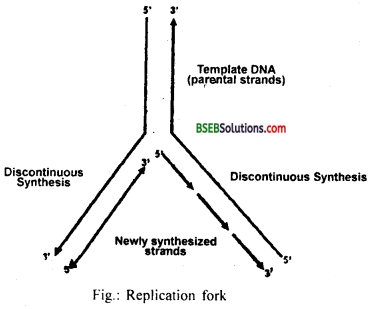
Question 13.
What are introns and exons? What process ensures a linear arrangement of amino acids although the genes are discontinuous?
Answer:
The eukaryotic gene is split into coding and non-coding sequences. The coding sequences also called as exons are those which are expressed in the mature RNA. They are flanked by non-coding sequences, termed Introns or intervening sequences which do not appear in mature or processed RNA. Splicing is the process which removes the unwanted mRNA regions and joins the functional regions responsible for coding.
![]()
Long Answer Type Questions
Question 1.
Discuss the mechanism of protein synthesis.
Answer:
Translation (protein synthesis) takes place in several steps :
(i) Activation of amino acids: It is carried out by aminoacyl tRNA synthetases. In the presence of ATP and mg:\ an amino acid combines with specific aminoacyl t-RNA synthetase. It produces aminoacyl-adenylate-enzyme complex.
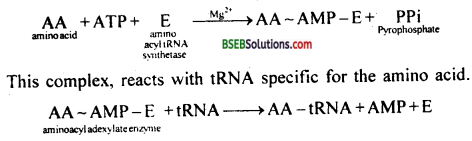
(ii) Initiation : It requires initiation factors. Procaryotes have three initiation factors IF3, IF2, and IF1. Eukaryotes have nine – eIF2, eIF3. elF1, eIF4, eIF4B, eIF4C, eIF4D, eIF5, eIF6.
For initiation, the ribosome binds to the mRNA at start codon AUG, which is recognised by initiator tRNA for methionine. It takes place in following steps :

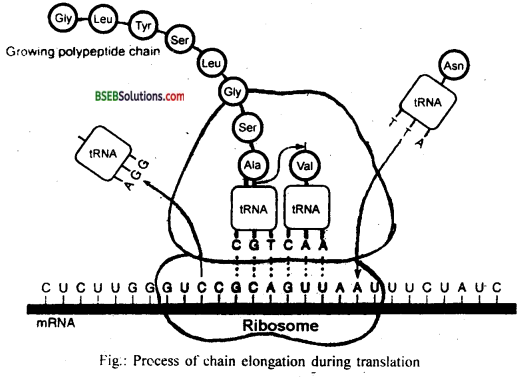
(iii) Elongation: During this phase, complexes composed of an amino acid linked to tRNA, sequentially bind to the appropriate codon in mRNA by forming complementary base pairs with the tRNA anticodon. The ribosome moves from codon to codon along with the mRNA. Amino acids are added one by one, translated into polypeptide sequences dictated by DNA and represented by mRNA. A peptide bond (-CO – NH-) is established between the carboxyl group (-COOH) of amino acid attached to tRNA at P-site and amino group (-NH) of amino acid attached to tRNA at A-site. Peptidyl transferase enzyme catalyses the formation of peptide bond.
For every single amino acid incorporated in peptide chain, l ATP and 2 GTP molecules are used.
(iv) Termination: Polypeptide synthesis terminates when a non-sense codon of mRNA reaches A-site. There are three non-sense codons – UAA, UAG and UGA. These are not recognised by any tRNA. The polypeptide is released in the presence of release factor on encounter of any non-sense codons.
![]()
Question 2.
(i) Briefly describe the process of transcription.
(ii) Distinguish amongst structural gene, regulator gene and operator gene.
Answer:
(i) Transcription is the formation of RNA over DNA template. It requires enzyme RNA polymerase and takes place in several steps :
(a) Activation of ribonucleotides: All the four types of nucleotides are activated through phosphorylation. Enzyme phosphorylase and energy is required for the process.
(b) DNA template: Each DNA segment has a promoter region and a terminator region. A promoter region has RNA polymerase recognition site. RNA polymerase binds at promoter site, unwind the DNA helix with the help of helicases and gyrases. One strand act as template strand. Transcription occurs at 5’→3′ direction.
(c) Base pairing: Ribonucleoside triphosphates form complementary pairs, with the nitrogen bases of DNA template using pyrophosphatase enzyme and releasing energy. U pairs with A; A with T, C with G and G with C.
![]()
Ribonucleotide + PPi + Energy
(d) Chain formation: With the help of RNA polymerase the adjacent ribonucleotides join.to form RNA chain. The rate of chain formation is 30 nucleotides per second. RNA synthesis stops with polymerase reaches terminator region.
(e) Separation of RNA: Terminator factor of Rho(P) factor has ATP-ase activity, it helps in the release of completed RNA chain.
(ii) Differences amongst regulator gene, operator gene, promoter gene and structural gene :
Regulator Gene Operator Gene Promoter Gene Structural Gene
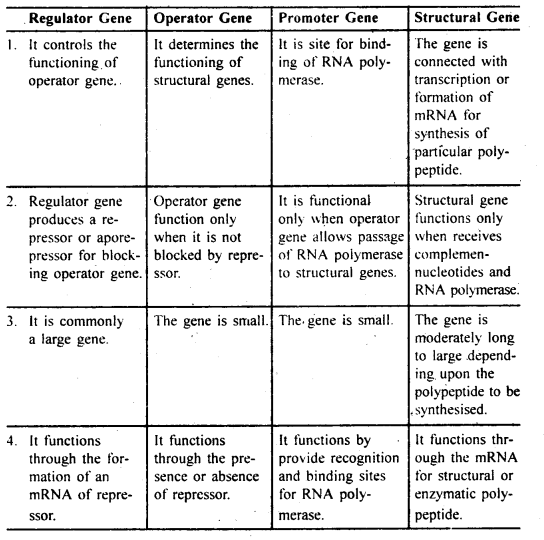
One Word Type Questions
Question 1.
Who coined the word gene?
Answer:
Johannsen.
Question 2.
Write any two non-sense codons.
Answer:
UGA, UAA.
Question 3.
How many bases code for one amino acid?
Answer:
Three.
Question 4.
Name any organism where RNA acts as genetic material.
Answer:
TMV.
![]()
Question 5.
Who first deciphered the genetic code?
Answer:
Nirenberg Ochoa. Hargobind Khorana and Francis Crick.
Question 6.
A nucleosome contains how many base pairs of DNA helix?
Answer:
200 base pairs.
Question 7.
What is genetic material?
Answer:
DNA.
Question 8.
Who suggested inborn errors in metabolism?
Answer:
Archibold Garrod.
Question 9.
mRNA of eukaryotes is synthesized with the help of what?
Answer:
RNA polymerase II.
Question 10.
Who proved experimentally that DNA replication is semi-conservative?
Answer:
Nesselson and Stahl.